Pre-print: Geospatial models of mutations encoding antimalarial resistance
Hot off the presses, here’s a new pre-print of spatiotemporal models of mutations in the malaria parasite associated with drug resistance. Artemisinin combination therapies are the most widely-used treatment for P. falciparum malaria, but the spread of artemisinin partial resistance into Africa raises the spectre of no effective anti-malarial treatment for populations with the greatest burden of the disease.
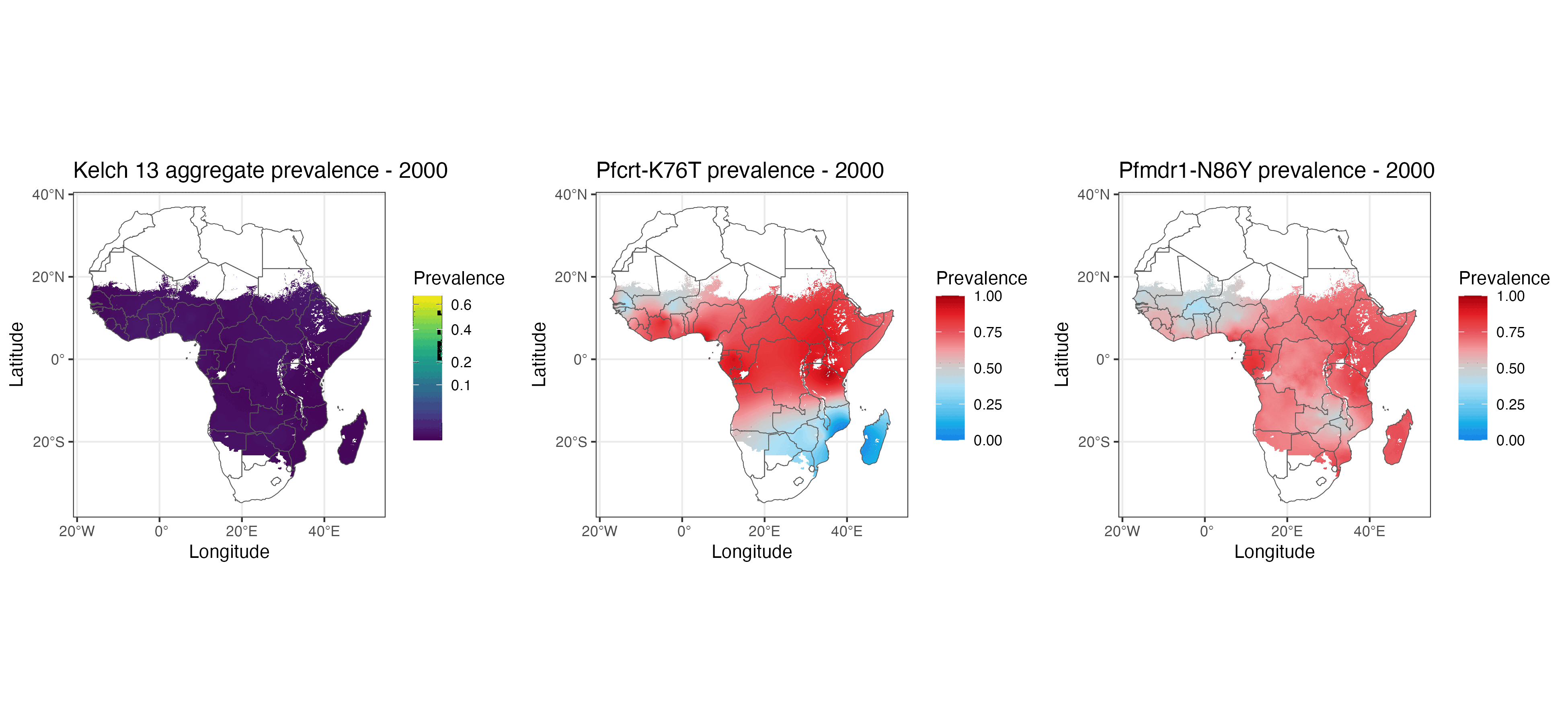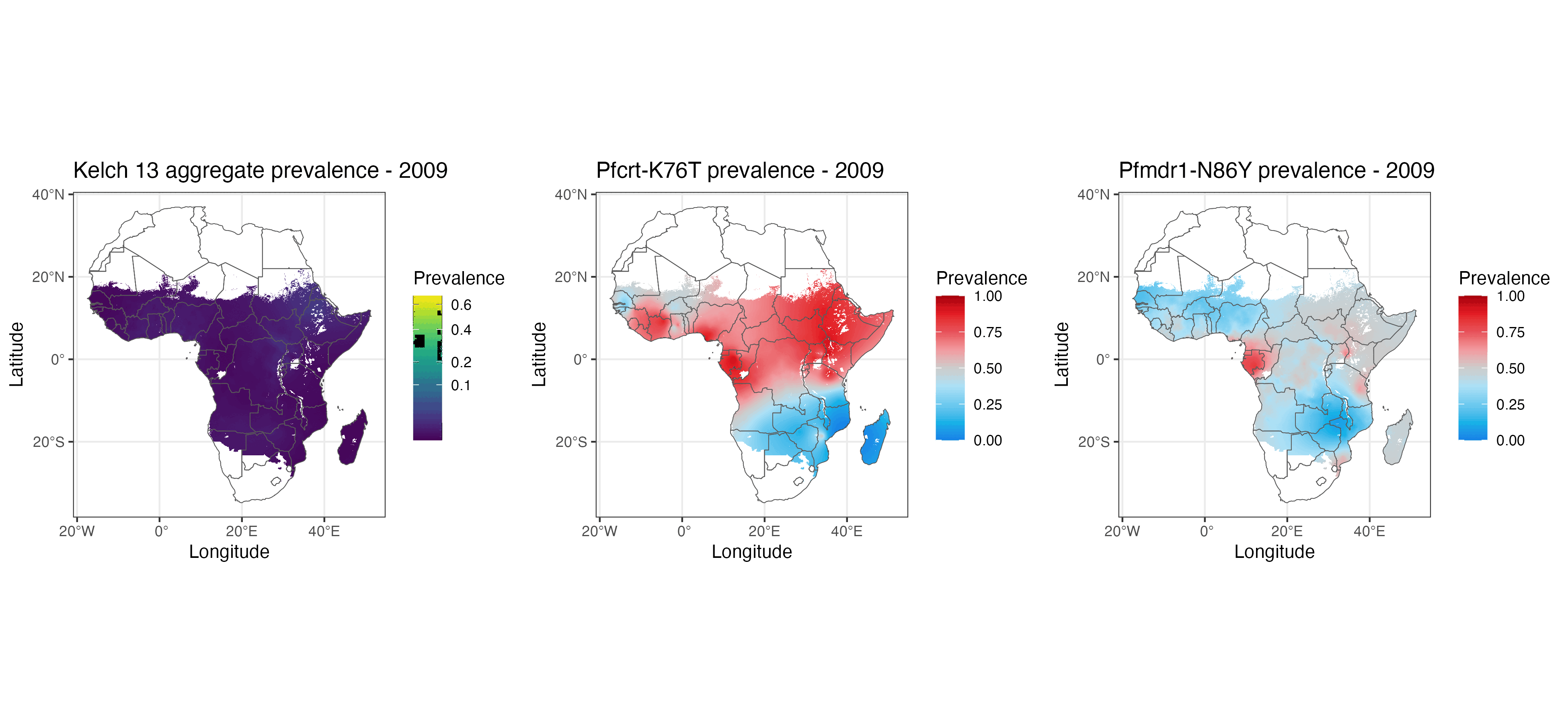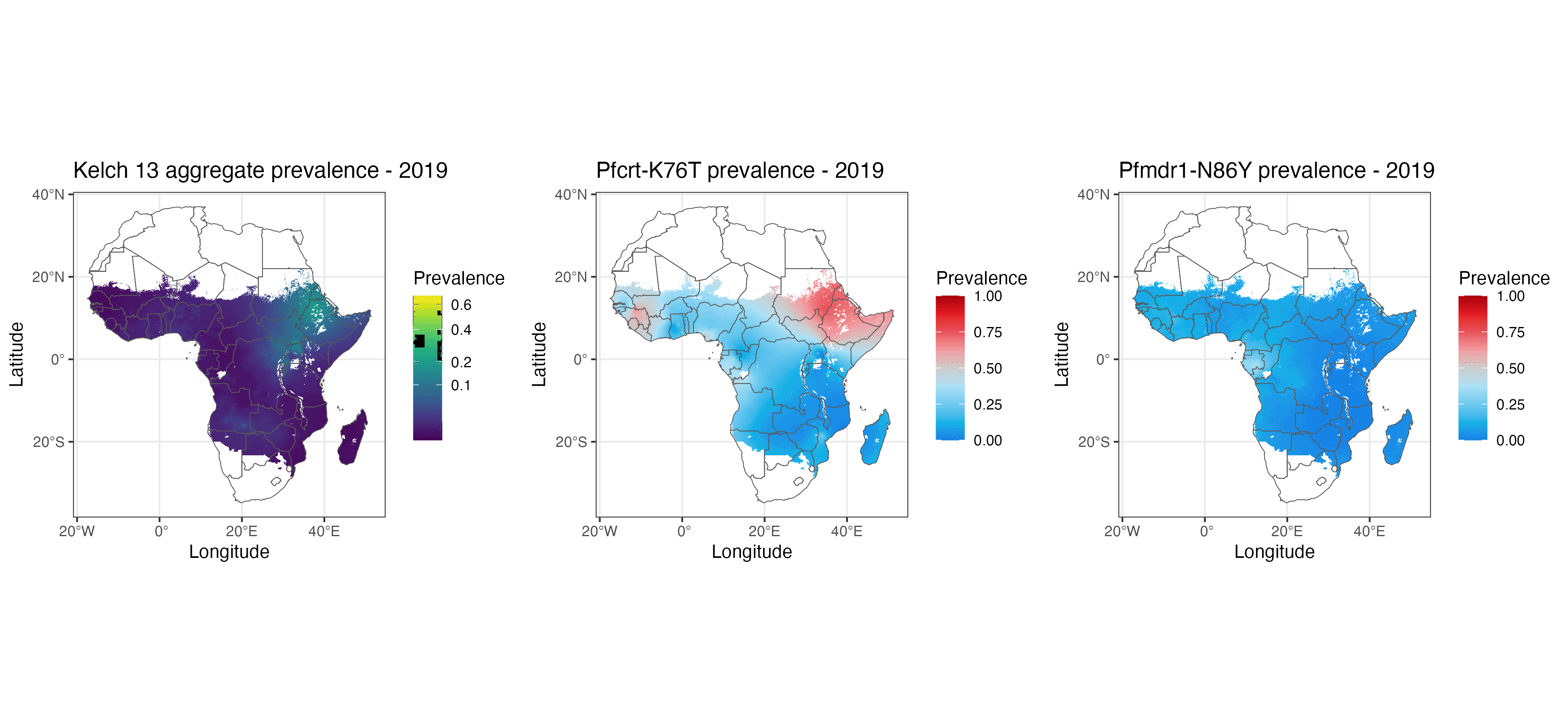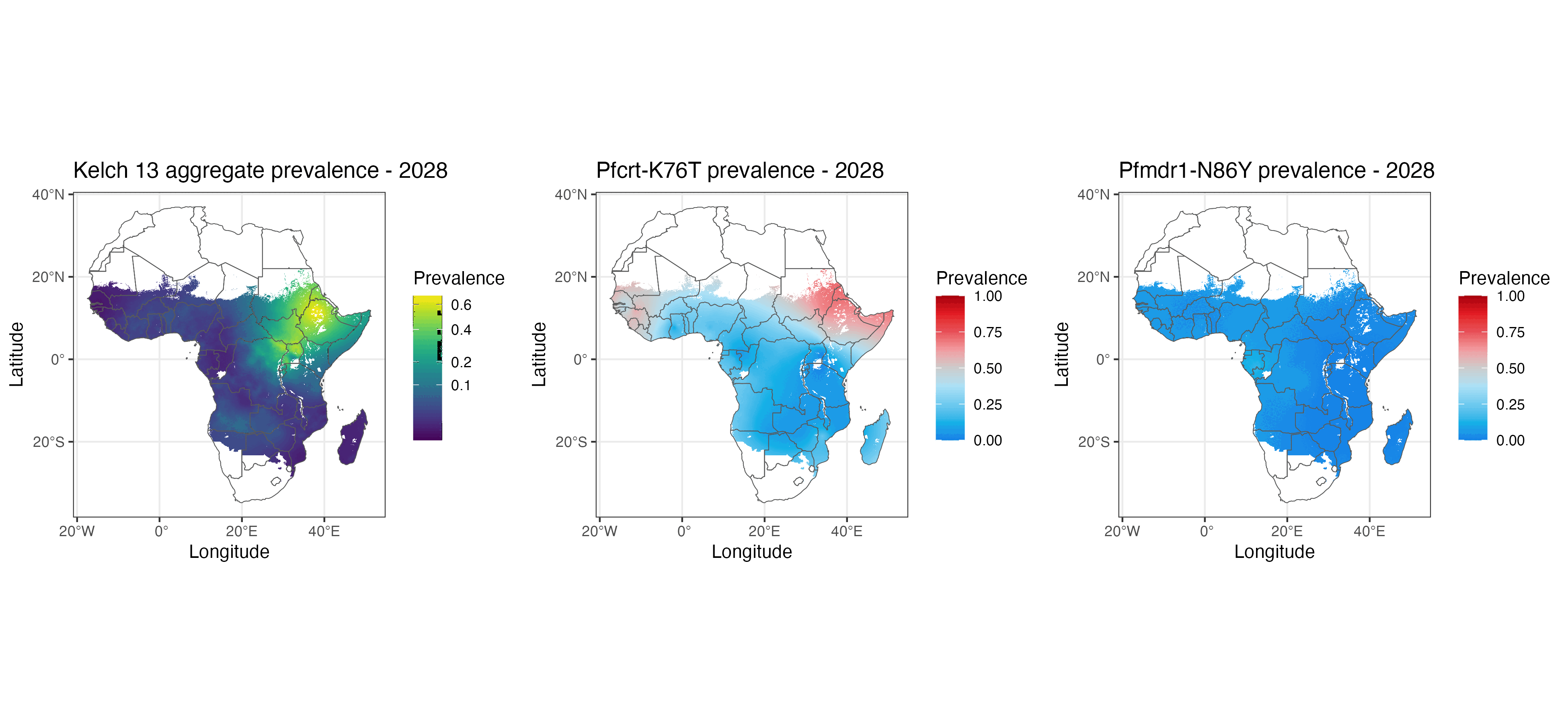
We present geostatistical models of mutations associated with malarial parasite resistance/reduced antimalarial drug susceptibility. These statistical models (1) allow us to visualise variation in observed marker prevalences, (2) allow us to extrapolate marker prevalence to areas in space and time that are not represented in surveillance data, and (3) identify geographical gaps to guide prospective sampling collection. The methods we describe can be used for many other datasets with incomplete data sampling, such as antimicrobial resistance as well as human and animal genetics.
This paper is the first big project of my postdoc with IDDO! Very exciting to have it submitted! Have a look at it here, or go straight to GitHub for the code.